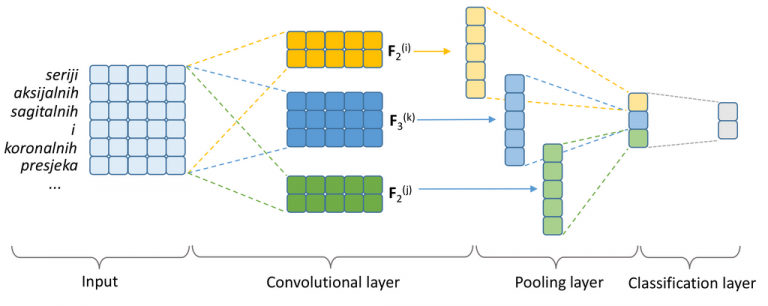
Narrative texts in electronic health records can be efficiently utilized for building decision support systems in the clinic, only if they are correctly interpreted automatically in accordance with a specified standard. This paper tackles the problem of developing an automated method of labeling free-form radiology reports, as a precursor for building query-capable report databases in hospitals. The analyzed dataset consists of 1295 radiology reports concerning the condition of a knee, retrospectively gathered at the Clinical Hospital Centre Rijeka, Croatia. Reports were manually labeled with one or more labels from a set of 10 most commonly occurring clinical conditions. After primary preprocessing of the texts, two sets of text classification methods were compared: (1) traditional classification models—Naive Bayes (NB), Logistic Regression (LR), Support Vector Machine (SVM), and Random Forests (RF)—coupled with Bag-of-Words (BoW) features (i.e., symbolic text representation) and (2) Convolutional Neural Network (CNN) coupled with dense word vectors (i.e., word embeddings as a semantic text representation) as input features. We resorted to nested 10-fold cross-validation to evaluate the performance of competing methods using accuracy, precision, recall, and F1 score. The CNN with semantic word representations as input yielded the overall best performance, having a micro-averaged F1 score of 86.7%. The CNN classifier yielded particularly encouraging results for the most represented conditions: degenerative disease (95.9%), arthrosis (93.3%), and injury (89.2%). As a data-hungry deep learning model, the CNN, however, performed notably worse than the competing models on underrepresented classes with fewer training instances such as multicausal disease or metabolic disease. LR, RF, and SVM performed comparably well, with the obtained micro-averaged F1 scores of 84.6%, 82.2%, and 82.1%, respectively.